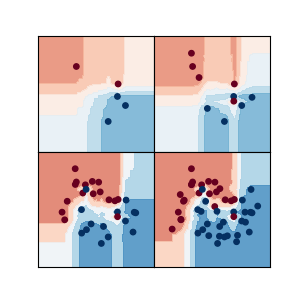
Classification¶
For now, onelearn provides mainly the AMFClassifier class for multi-class
classification.
Its usage follows the scikit-learn API, namely a partial_fit, predict_proba
and predict methods to respectively fit, predict class probabilities and labels.
The AMFClassifier with default parameters is created using
from onelearn import AMFClassifier
amf = AMFClassifier(n_classes=2)
where n_classes must be provided for the construction of the object.
Also, a baseline dummy classifier is provided by OnlineDummyClassifier, see below.
onelearn.AMFClassifier(n_classes[, …]) |
Aggregated Mondrian Forest classifier for online learning. |
onelearn.OnlineDummyClassifier(n_classes[, …]) |
A dummy online classifier only using past frequencies of the labels. |
About AMFClassifier¶
The AMFClassifier class implements the “Aggregated Mondrian Forest” classifier
for online learning (see reference below). This algorithm is truly online, in the sense
that a single pass is performed, and that predictions can be produced anytime.
For multi-class classification with \(C\) classes, we observe, before time \(t\), pairs of features and labels \((x_1, y_1), \ldots, (x_{t-1}, y_{t-1})\) where \(y_s \in \{ 1, \ldots, C \}\) for each \(s = 1, \ldots, t-1\).
Each node in a tree predicts according to the distribution of the labels
it contains. This distribution is regularized using a Dirichlet (a.k.a “Jeffreys”) prior
with parameter \(\alpha > 0\) which corresponds to the dirichlet parameter in AMFClassifier.
Each node \(\mathbf v\) of a tree predicts, before time \(t\), the probability of
class \(c\) as
for any \(c = 1, \ldots, C\), where \(n_{{\mathbf v}, t}(c)\) is the number of samples of class \(c\) in node \(\mathbf v\) before time \(t\). This formula is therefore simply a regularized version of the class frequencies.
Each node \(\mathbf v\) in the tree corresponds to a cell denoted \(\mathrm{cell}({\mathbf v})\), which corresponds to a hyper-rectangular subset of the features space. The predictions of a node, before time \(t\), are evaluated by computing its cumulative loss as
which is the sum of the prediction losses of all the samples whose features belong to
\(\mathrm{cell}({\mathbf v})\).
By default, we consider, for multi-class classification, the logarithmic loss
\(\ell (\widehat y, y) = - \log (\widehat y(y))\) for \(y \in \{ 1, \ldots, C \}\).
The loss can be changed using the loss parameter from AMFClassifier (however
only loss="log" is supported for now).
Given a vector of features \(x\) and any subtree \(\mathcal T\) of the current tree, we define \(\mathbf v_{\mathcal T}(x)\) as the leaf of \(\mathcal T\) containing \(x\) (namely \(x\) belongs to its cell). The prediction at time \(t\) of the subtree \(\mathcal T\) for \(x\) is given by
namely the prediction of \(\mathcal T\) is simply the prediction of the leaf of \(\mathcal T\) containing \(x\). We define also the cumulative loss of a subtree \(\mathcal T\) at time \(t\) as
When use_aggregation is True (the highly recommended default), the prediction function
of a single tree in AMFClassifier is given, at step \(t\), by
where the sum is over all subtrees \(\mathcal T\) of the current tree, and where the prior \(\pi\) on subtrees is the probability distribution defined by
where \(|\mathcal T|\) is the number of nodes in \(\mathcal T\) and \(\eta > 0\)
is the learning rate that can be tuned using the step parameter in AMFClassifier
(theoretically, the default value step=1.0 is the best, and usually performs just fine).
Note that \(\pi\) is the distribution of the branching process with branching probability \(1 / 2\) at each node of the complete binary tree, with exactly two children when it branches. This aggregation procedure is a non-greedy way to prune trees: the weights do not depend only on the quality of one single split but rather on the performance of each subsequent split.
The computation of \(\widehat {f_t}(x)\) can seem computationally infeasible, since it involves a sum over all possible subtrees of the current tree, which is exponentially large. Besides, it requires to keep in memory one weight \(e^{-\eta L_{t-1} (\mathcal T)}\) for all the subtrees \(\mathcal T\), which seems exponentially prohibitive as well !
This is precisely where the magics of AMFClassifier resides: it turns out that
we can compute exactly and very efficiently \(\widehat {f_t}(x)\) thanks to the
prior choice \(\pi\) together with an adaptation of the Context Tree Weighting algorithm,
for which more technical details are provided in the paper cited below.
The interested reader can find also, in the paper cited below, the construction details of
the online tree construction, which is based on the Mondrian process and Mondrian Forests.
Finally, we use \(M\) trees in the forest, all of them follow the same randomized construction. The predictions, for a vector \(x\), of each tree \(m = 1, \ldots, M\), are denoted \(\widehat {f_t}^{(m)}(x)\). The prediction of the forest is simply the average given by
The number of trees \(M\) in the forest can be tuned with the n_estimators parameter
from AMFClassifier, the default value is 10, but the larger the better
(but requires more computations and memory).
Note
When creating a classifier instance, such as a AMFClassifier object, the
number n_classes of classes must be provided to the constructor.
Note
All the parameters of AMFClassifier become read-only after the first call
to partial_fit
References¶
@article{mourtada2019amf,
title={AMF: Aggregated Mondrian Forests for Online Learning},
author={Mourtada, Jaouad and Ga{\"\i}ffas, St{\'e}phane and Scornet, Erwan},
journal={arXiv preprint arXiv:1906.10529},
year={2019}
}